NEWS

Yong Cang, professor of ShanghaiTech University and co-founder of Degron Therapeutics
The Challenge
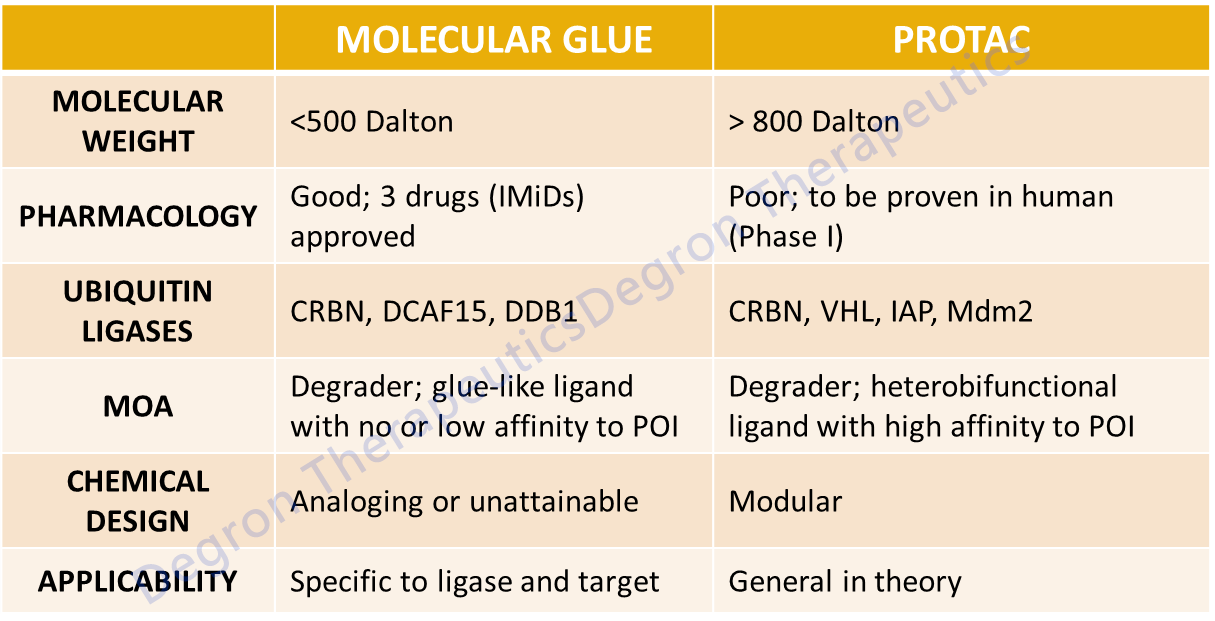
Targeted protein degradation with small molecule compounds is an emerging strategy to inactivate therapeutic targets that are intractable by traditional pharmacological approaches1,2. Heterobifunctional PROTACs (proteolysis targeting chimeras) ligand targets to ubiquitin ligases, but the molecular weight and pharmaceutical properties of PROTACs fall outside of the typical range of chemical drugs3. By contrast, much smaller molecular glue degraders, including thalidomide analogues4-7 and aryl sulfonamides8-10, induce complementary protein-protein interactions between cullin 4-RING E3 ubiquitin ligase (CRL4) and various targets, leading to target ubiquitination and depletion. However, all these molecular glue degraders have thus far only been found serendipitously, and systematic discovery of novel molecular glue degraders is not yet in place.
The Paper
On June 3 this year, Benjamin Ebert and Nicolas Thoma published in Nature the first undertaking to proactively discover molecular glue degraders11. By correlating compound cytotoxicity and ubiquin ligase expression across hundreds of cancer cell lines, they identified the CDK12 kinase inhibitor CR8 as a bona fide glue molecule that bridges the interaction between CDK12 and DDB1, the CRL4 adaptor, thus exposing CDK12-bound cyclin K to the range for modification by CRL4. In this Nature Chemical Biology paper12, George Winter identified different chemotypes as molecular glue degraders of cyclin K, following the same mechanism of action as published earlier. However, Winter took a totally different approach; he started with a phenotypic screen for compounds that could kill cancer cells in a way dependent on cellular cullin ubiquitin ligase activities. This screen format can be applied to compound libraries beyond the 2000 cytotoxic chemicals used by Winter or the preclinical and clinical compounds covered by Ebert and Thoma, for discovery of novel degraders with novel mechanisms. Finally, the cyclin K degraders from both studies harbor a pyridyl moiety that induces the CDK12-DDB1 complex formation. It might be interesting to explore this chemical space to rationally design DDB1-targeted glue molecules.
My Research
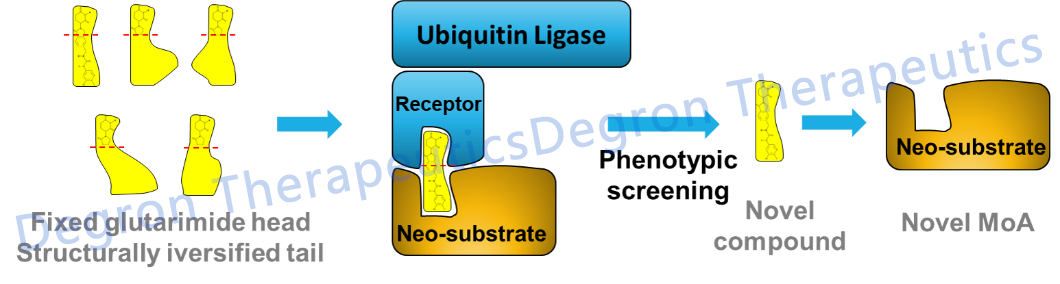
My lab has been working on the biology of CRL4 ubiquitin ligases in the past 13 years, always with an eye on targeting CRL4 for therapeutic development. After moving to the vibrant and resourceful ShanghaiTech University 3 years ago, my lab took a bold chemical biology approach to discover molecular glue degraders of oncoproteins. Our workis based on a hypothesis that chemical alteration of the surface of CRBN, a CRL4 substrate receptor, would enable binding to neo-substrates. By combining phenotypic screen and proteomic analysis, we have identified novel chemotypes that deplete two known oncology targets that regulate the cell cycle checkpoint and KRAS signaling respectively.
The Translation
Criteria to evaluate the impact of any biomedical research must include its long term benefits to human health. Several biotech startups founded by academic researchers are already dedicating themselves to translating lab discoveries to clinical development. Monte RosaTherapeutics in Viena and Coho Therapeutics in San Diego employ structural insights into substrate-ligase interactions to screen for novel glue molecules. Degron Therapeutics in Shanghai designs glue molecule libraries and develop cancer therapeutics with phenotypic screen and target discovery. Vividion Therapeutics and BioTheryX, both in San Diego, couple targetable ubiquitin ligases with molecular glue screens, as a backup to their PROTAC pipelines. Bigpharmas including Novartis and BMS have internal targeted protein degradation programs in which molecular glue discovery is an integral part.
In Summary, molecular glue degraders are clinically proven and more drug-like than PROTACs, but at this stage not amenable to chemical design if specific targets of interest are to be degraded. However, collective efforts from academia and industry are on track to unravel increasing number of glue structures together their targets, eventually allowing us to rationally design such molecules.
It took 6 years for Arvinas to bring the first PROTAC to be tested in the first patient. We should be optimistic that molecular glue degrader discovery might break the design bottleneck and expand the approved drugs in this category much sooner than we expected.
References:
1.Schapira, M., Calabrese, M. F., Bullock,A. N. & Crews, C. M. Targeted protein degradation: expanding the toolbox.Nat Rev Drug Discov 18, 949-963, doi:10.1038/s41573-019-0047-y (2019).
2.Chopra, R., Sadok, A. & Collins, I. A critical evaluation of the approaches to targeted protein degradation for drug discovery. Drug Discov Today Technol 31, 5-13, doi:10.1016/j.ddtec.2019.02.002 (2019).
3.Nguyen, K. M. & Busino, L. Targeting the E3 ubiquitin ligases DCAF15 and cereblon for cancer therapy. Semin Cancer Biol https://doi.org/10.1016/j.semcancer.2020.03.007,doi:10.1016/j.semcancer.2020.03.007 (2020).
4.Matyskiela, M. E. et al. A novel cereblon modulator recruits GSPT1 to the CRL4(CRBN) ubiquitin ligase. Nature 535, 252-257,doi:10.1038/nature18611 (2016).
5.Petzold, G., Fischer, E. S. & Thoma, N. H. Structural basis of lenalidomide-induced CK1alpha degradation by the CRL4(CRBN) ubiquitin ligase. Nature 532, 127-130, doi:10.1038/nature16979 (2016).
6.Kronke, J. et al. Lenalidomide causes selective degradation ofIKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301-305,doi:10.1126/science.1244851 (2014).
7.Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305-309,doi:10.1126/science.1244917 (2014).
8.Bussiere, D. E. et al. Structural basis of indisulam-mediated RBM39 recruitment to DCAF15 E3 ligase complex. Nat Chem Biol 16, 15-23,doi:10.1038/s41589-019-0411-6 (2020).
9.Uehara, T. et al. Selective degradation of splicing factor CAPERalpha by anticancer sulfonamides. Nat Chem Biol 13, 675-680,doi:10.1038/nchembio.2363 (2017).
10.Han, T. et al. Anticancer sulfonamides target splicing by inducing RBM39 degradation via recruitment to DCAF15. Science 356,doi:10.1126/science.aal3755 (2017).
11.Slabicki, M. et al. The CDK inhibitor CR8 acts as a molecular glue degrader that depletes cyclin K. Naturehttps://doi.org/10.1038/s41586-020-2374-x, doi:10.1038/s41586-020-2374-x(2020).
12.Mayor-Ruiz, C. et al. Rational discovery of molecular glue degraders via scalable chemical profiling. Nat Chem Biol,doi:10.1038/s41589-020-0594-x (2020).

